Building an analytics platform using the OMOP common data model
Building an analytics platform using the OMOP common data model
With the digital transformation of healthcare information and the shift towards value-based care (sometimes called accountable care), big data has become increasingly relevant to the healthcare industry.
In healthcare, big data relates to large quantities of data, often obtained from various sources and in varying formats. This makes the data dense and complex, meaning it requires sophisticated methods to collect and analyze.
Where healthcare organizations can invest in big data, they could make significant improvements for patients, particularly in:
- Preventive care
- Medication error rates
- Disease control
Using data from several sources
Hospitals and other healthcare organizations use numerous databases to store information. These range from patient-focused databases like Electronic medical records (EMR) and Electronic health records (EHR), to business-focused databases for finances and costs.
While the disparate pools of information contained within these databases present fertile ground for analysis, the information is usually stored in non-standardized formats that can’t be compared and contrasted.
To make the information useful, it must be viewed within an effective analytical framework.
What is a common data model?
A common data model (CDM) is a shared language that applies syntactic and semantic consistency to observational data obtained from multiple sources. A CDM facilitates interoperability by transforming data into unambiguous, exchangeable information that can be read by computers and humans. This can help organizations align data within their own departments, as well as exchange data with other organizations.
In the illustration, you can see a representation of this process as applied with the Observational Medical Outcomes Partnership (OMOP) CDM.
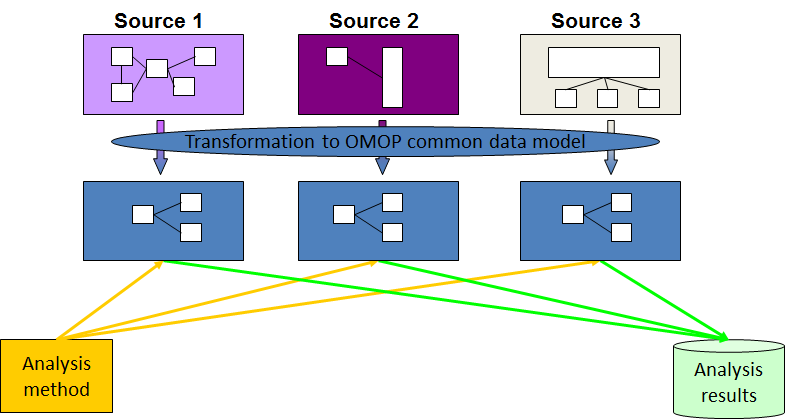
Source: https://www.ohdsi.org/data-standardization/the-common-data-model/
Raw data from various sources, each with their own attributes and relationships, is mapped to the OMOP CDM. Through this process, the data becomes standardized so it can be effectively exchanged and analyzed.
What is the OMOP CDM?
The OMOP CDM is a healthcare research-focused data model maintained by the Observational Health Data Sciences and Informatics initiative (OHDSI).
Though the OMOP CDM was originally centered around drug safety, it has since expanded to include other analytical use cases such as comparative effectiveness of medical interventions and health system policies (you can read more about how OHDSI is developing the OMOP CDM here).
OMOP has designed a CDM that is optimized for typical observational research purposes. As OHDSI writes on GitHub, these are:
- Identifying patient populations with certain healthcare interventions and outcomes
- Characterizing these patient populations for various parameters
- Predicting the occurrence of these outcomes in individual patients
- Estimating the effect these interventions have on a population
In the illustration you can see an overview of all the tables found in the OMOP CDM.
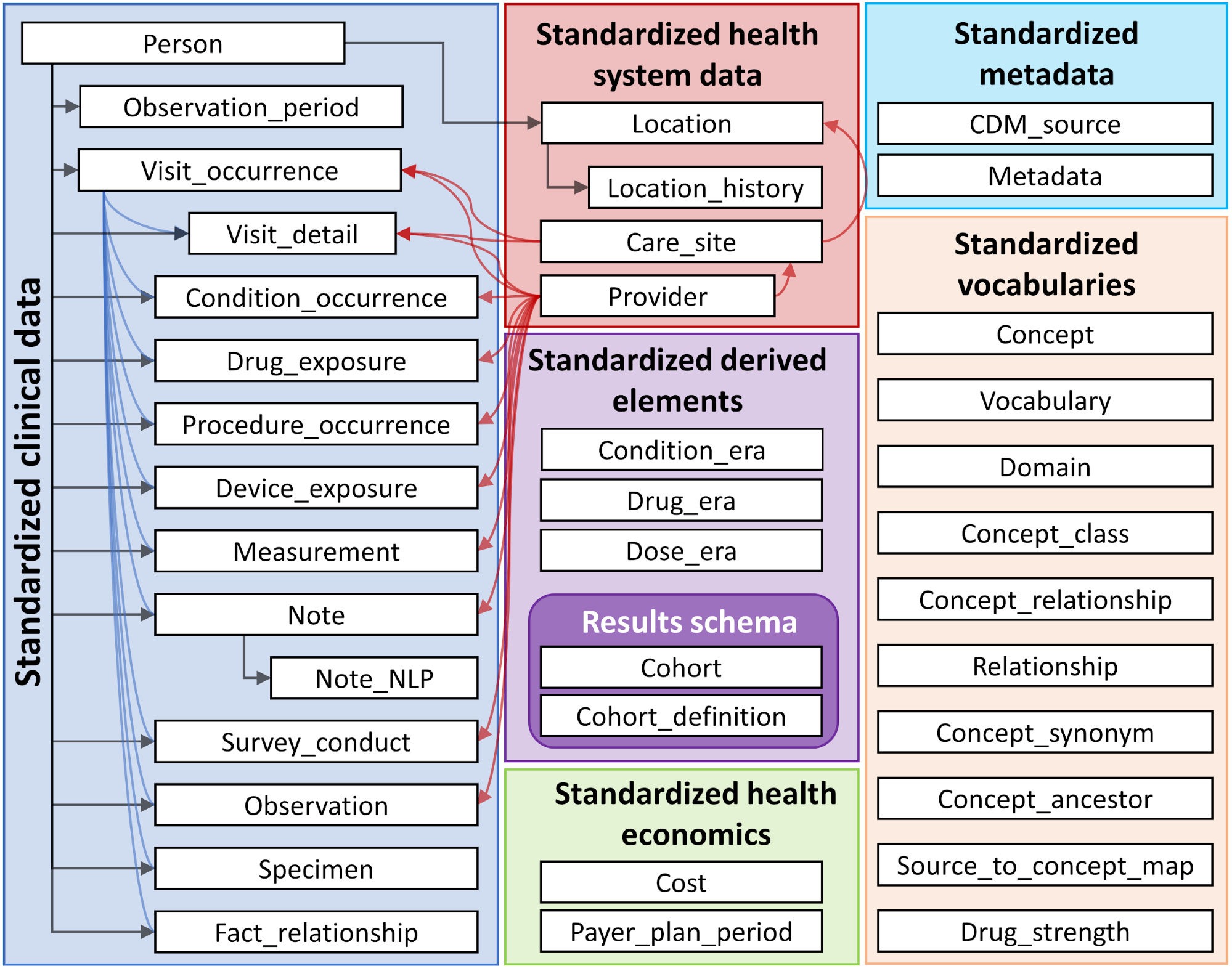
Source: https://ohdsi.github.io/TheBookOfOhdsi/CommonDataModel.html
The following list presents some of the design concepts used in the OMOP CDM schema:
- All clinical event tables are linked to the PERSON table
- Data that can be used to identify patients is very limited
- Separation between read-only and read-write tables – tables that need to be manipulated by web-based tools or end users are stored in the RESULTS schema
- Each Standard Concept has a unique Domain assignment, which defines which table they are recorded in (like drug, procedure, condition, observation, and so on)
- Variable names across all tables follow one convention
Find more details about the OMOP CDM in the book of OHDSI - chapter 4.
The Data4Life Analytics Platform
Data4Life provides a health platform with a global focus that acts as a service-enabler for patients, healthcare providers, and researchers. It allows its partners to focus on their core competencies while handling sensitive health data in a safe and secure manner.
In collaboration with care providers, research institutions, and health tech companies, our goal is to build a rich collection of observational healthcare data that will be made available for clinical research purposes. Data4Life is a nonprofit organization and we hope to facilitate this research to improve people’s wellbeing.
The data ingested into our Analytics Platform (ALP) will be accessed under specific governance rules for conducting a research study. By providing a platform to connect researchers with healthcare data, Data4Life will enable the discovery and validation of new insights faster than was ever possible before.
Data4Life builds the platform on top of the OMOP CDM as the data model to store healthcare data. Utilizing the OMOP CDM, Data4Life can build on top of the open source tools provided by the OHDSI community and contribute to the community by extending the available tools.
Analytics tools using OMOP CDM
Since data analysis requires flexibility, it’s important to enable researchers with a technical background to analyze the data through advanced programming languages, like Python or R. In a second phase, once typical usage patterns are well understood, additional UI support might be added for less technical users.
The following illustration shows the architecture of the analytical platform developed by Data4Life.
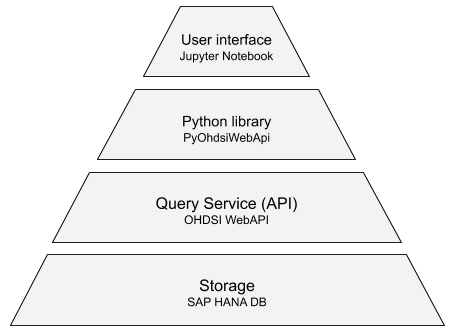
Storage
The database system is one of the most performance critical components. To ensure fast analytics performance that can enable an interactive user experience, Data4Life plans to store observational healthcare data on SAP HANA, a columnar database that can compress and store large amounts of data in-memory. As mentioned before, the schema selected for storing the data is the OMOP CDM.
Query Service (API)
Data4Life is using the OHDSI WebAPI as a layer to interact with the databases available on the platform. OHDSI WebAPI, a middleware offered by OHDSI, fits the bill perfectly as it translates criteria to a SQL query executable on an OMOP database and enforces security.
OHDSI WebAPI has been modified by Data4Life to enable integration with the HANA database.
Python query library
Machine learning has become very important in today’s research and Python has undoubtedly gained popularity in this space. Hence, at Data4Life, we have developed a library (PyOhdsiWebApi) that allows the creation of patient cohorts directly in Python, making them available for further analysis using analytic Python libraries such as NumPy, PyTorch, and others.
The key objective of the library is to write complex criteria in a simple query language rather than with SQL, because SQL statements can become complicated and prone to errors as the criteria grows. PyOhdsiWebApi interfaces with WebAPI by consuming its RESTful services.
We will continue to enhance the library and add more features on the basis of the feedback given by the researchers using this library. Data4Life is in the process of making the Python library an open-source code as a contribution to the OHDSI community.
Jupyter Notebook
The Jupyter Notebook is an open-source web application that allows data scientists to create and share documents that contain:
- Live code
- Equations
- Visualizations
- Narrative text
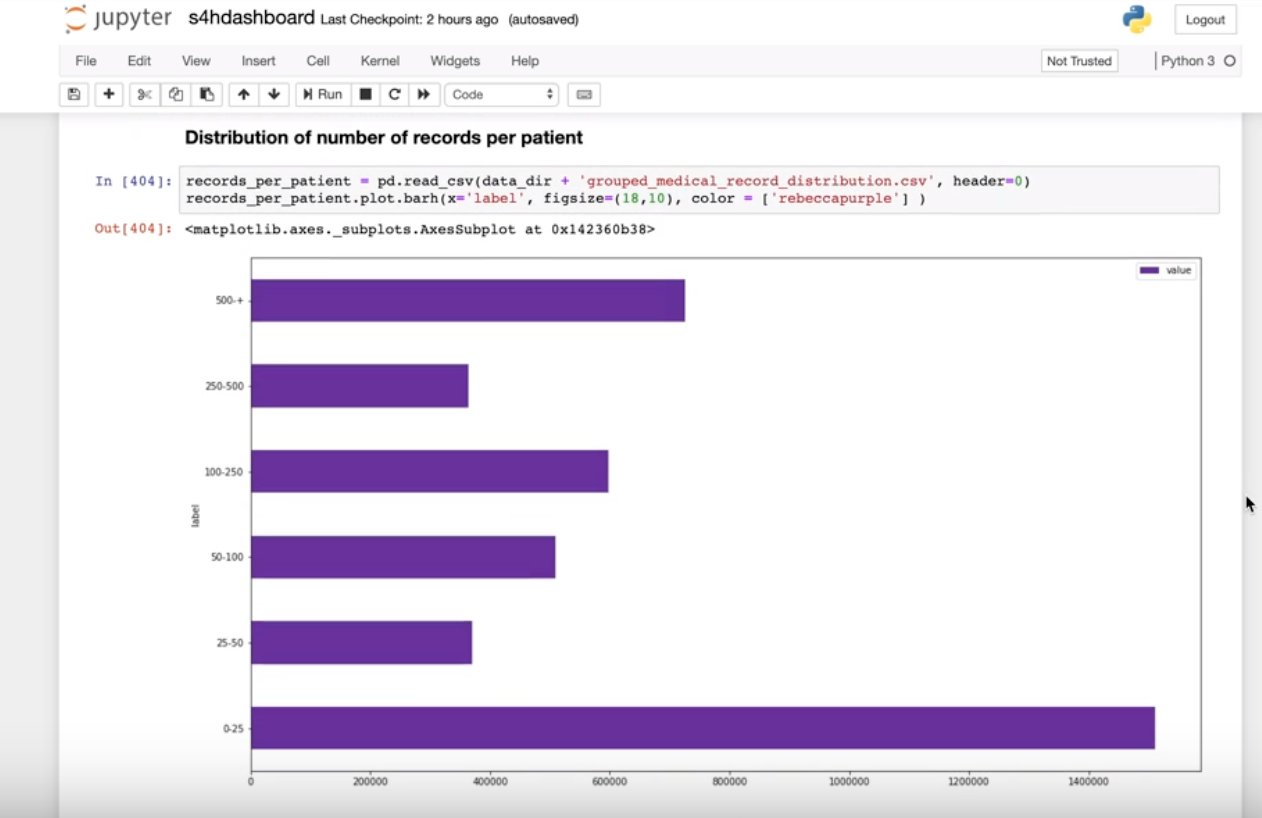
Data4Life integrated Jupyter Notebook to work seamlessly with the Python query library. With Jupyter Notebook data, scientists can build their cohorts, analyze them (including applying advanced machine or deep learning methods), and visualize the results.
What’s next?
Thanks to the OMOP CDM and the OHDSI community initiatives, Data4Life is closer to realizing its vision of connecting researchers and healthcare data to improve global health.
Find out more about what we do on the Data4Life website and connect with us on LinkedIn to stay up-to-date with our journey.
Related articles
Leveraging the OHDSI OMOP common data model for research using observational health data